Protein Target Deconvolution & Network Pharmacology
A semi-automated computational pipeline utilizing reverse docking and PPI network analysis to unveil the mechanisms of uncharacterized cytotoxic compounds.
The Bottleneck in Natural Product Discovery A persistent challenge in pharmacognosy and synthetic organic chemistry is the identification of highly active, newly extracted or synthesized compounds whose exact biological targets remain unknown. Wet-lab cell viability assays (e.g., testing against HL-60 cell lines) prove cytotoxicity, but without a Mechanism of Action (MoA), these findings often stall in lower-quartile journals.
The IDD Lab Solution: A Semi-Automated Deconvolution Pipeline We have engineered a robust, end-to-end computational pipeline designed to map the therapeutic mechanisms of uncharacterized compounds. By integrating high-throughput reverse docking with systems biology, we transition research from simple phenotypic observation to target-specific validation.
Methodology & Workflow:
- High-Throughput Reverse Docking: We screen the novel ligand against comprehensive protein target databases to identify high-affinity binding candidates.
- Protein-Protein Interaction (PPI) Profiling: Top-scoring targets are mapped into complex PPI networks to deduce the systems-level downstream effects and identify the most critical node (essential hits).
- In-Depth Binding Analysis: Rigorous rescoring and interaction mapping of the ligand within the essential hit’s active site.
- Molecular Dynamics (MD) Validation: Ensuring thermodynamic stability of the predicted complex over time to validate the structural hypothesis.
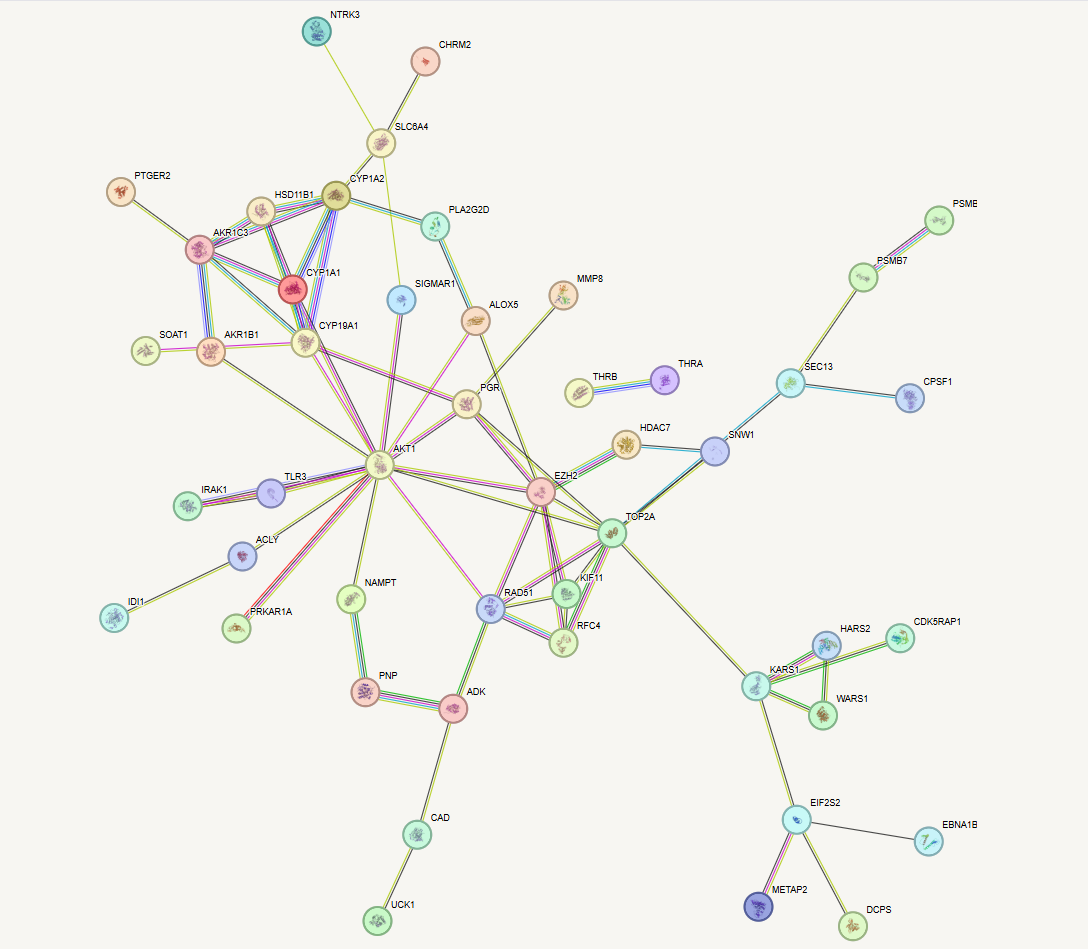
Proof of Concept & Collaboration This pipeline is currently validating the selective cytotoxicity of novel plant-extracted compounds against HL-60 cell lines.
Call for Collaboration: IDD Lab actively seeks partnerships with wet-lab investigators. We offer this pipeline to uncover the mechanisms of your novel compounds, elevating standard extraction papers into comprehensive, Q1-level mechanistic studies.
GitHub Repository: Semi-Automated Reverse Docking